Welcome to T-REX
Software for the processing and analysis of T-RFLP data
Overview T-RFLP data analysis requires many steps that are often time-intensive, error-prone, and can introduce unwanted variability into the analysis.
T-REX is a free, online tool that was developed to address these obstacles.
T-REX allows users to:
- Label raw data with attributes related the experimental design of the samples
- Determine a baseline threshold for identification of true peaks over noise
- Align T-RFs in all samples (bin T-RFs)
- Construct a two-way data matrix from labeled data and manipulate the matrix in a variety of ways
- Produce several measures of data matrix complexity, including the distribution of variance between main and interaction effects and sample heterogeneity
- Analyze a data matrix with the Additive Main Effects and Multiplicative Interaction Model (AMMI)
T-REX has been designed and built with flexibility in mind. Each of the menu buttons on the left correspond to a certain action performed on the data. Since these actions are to a large extent independent,
any of the buttons can be used at any time, without the need to reload or upload the same data again. Although no particular sequence of actions is imposed on the user, a typical flow of analysis is illustrated below.
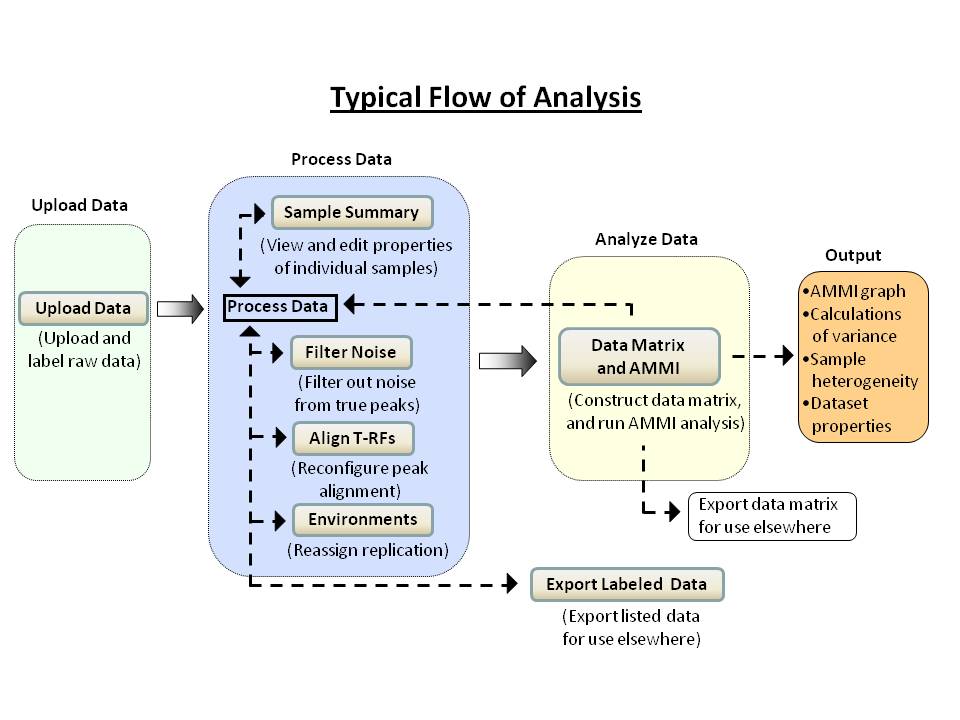
It is recommended that users read the program
documentation to become familiar with the functions and processes of this software before starting. An excerpt from the documentation, providing a bird's-eye view of the program funcitons is available
here. On-line help is also available, accessible from each of the sub-pages.
Getting started:
About T-REX:
T-REX was developed by Robert Bukowski under the guidance of Steve Culman, Hugh Gauch, Hinsby Cadillo-Quiroz and Dan Buckley.
Please cite the program as:
Culman, S.W., Bukowski, R., Gauch, H.G., Cadillo-Quiroz, H., Buckley, D.H. 2009.
T-REX: Software for the Processing and Analysis of T-RFLP data. BMC Bioinformatics 10:171
Full text of the article can be found
here T-REX was written using Microsoft ASP.NET and MS SQL Server platforms on Windows Server 2003. It is hosted at the Computational Biology Service Unit (CBSU) at Cornell University.
T-REX can be found at the web address: http://trex.biohpc.org/. The access to the program is free and requires only a web browser and an internet access.
In the future, the source code of
T-REX will be available under the GNU GPL License.
Funding for
T-REX's development was made possible by the National Science Foundation's Biogeochemistry and Biocomplexity IGERT grant, and by the Microsoft Corporation.
Please email any software-related technical questions to
Robert Bukowski and general comments or questions to
Steve Culman.